In a new study, researchers from the National Institute of Diabetes and Digestive Diseases, the National Cancer Institute, the Shake Institute for Biology, and the Scripps Research Institute have discovered a key part of how a powerful class of HIV drugs binds the HIV intasome. By solving for the first time the three-dimensional structure of this intasome when combined with different drugs, they discovered what makes this class of drugs so effective. This study provides important insights that may help to design or improve new therapies for HIV. The relevant findings were recently published in the journal Science, and the paper was entitled “Structural basis for strand transfer inhibitor binding to HIV intasomes”.
Dmitry Lyumkis, the corresponding author of the paper and assistant professor of genetics laboratory at the Shake Institute for Biological Research, said, “The drugs we studied are the latest compounds available in clinical practice today, with several important preclinical molecules. Until then, no one knows exactly how they bind to this HIV intasome. A better understanding of the effects of these drugs will help us to improve them and design new therapeutic compounds.”
HIV intasome is a key structure of this viral infection, which consists of HIV protein integrase and viral DNA strands, and are formed when this virus invades human cells. This intasome enters human cells and subsequently undergoes the necessary chemical reaction, thereby integrating the genetic material of this virus into human DNA.
Some drugs called integrase strand transfer inhibitor (INSTI) successfully block this intasome; when it cannot integrate viral DNA into the human genome, HIV is not able to infect human cells. Currently, four INSTI drugs are approved by the US Food and Drug Administration (FDA), and some are under development.
Despite the success of these molecules, efforts are still being made to investigate how to inhibit HIV intasome, mainly due to the difficulty in isolating such intasome for structural studies. In the past, most studies on HIV intasome and INSTIs were performed on another retrovirus called prototype foamy virus (PFV). In 2017, Lyumkis and colleagues first solved the structure of purified HIV intasome (Science, 2017, doi: 10.1126/science. Aah5163).
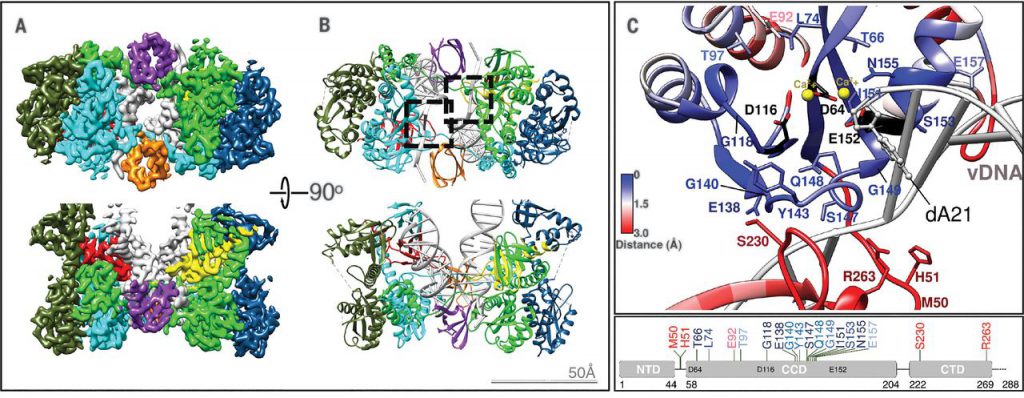
In this new study, Lyumkis’s team went further: they obtained the structure of the HIV intasome when blocked by one of four INSTIs—the commercially available drug bictegravir; and three experimental compounds called 4f, 4d, and 4c. They used single-particle cryo-electron microscopy (an imaging technique they helped optimize) to reveal the structure of each HIV intasome-drug complex.
The first observation of Lyumkis is that these drugs are different when combined with HIV intasome than when they are combined with PFV intasome. For example, compound 4f loops back to itself when bound to PFV intasome, but remains relatively flat when bound to HIV intasome, and these details can help improve the binding properties of potential future molecules.
Dario Oliveira Passos, the co-first author of the paper and a researcher in the Lyumkis laboratory, said, “So far, everyone is still using PFV intasome structures to rationally think and understand the mechanism of action of these drugs. But we found that if we want to make further progress, this field does need to move forward and study HIV intasome structure.”
Min Li, the co-first author of the paper and a researcher at the National Institute of Diabetes and Digestive Diseases, said, “We and many others have worked for decades to achieve the goal. Excitingly, we can finally understand the detailed working principles of such HIV inhibitors and help develop new drugs.”
These structures also reveal why these drugs are so effective and what makes them so outstanding in avoiding drug resistance. Lyumkis and colleagues found that these INSTIs fill the entire space usually occupied by DNA. This means that if HIV intasome undergoes mutations that prevent INSTI drug binding, it will also make this intasome unable to invade human cells.
Finally, these very high-resolution structures obtained by these researchers allowed them to observe detailed information about how these drugs interact chemically with this binding pocket and how these INSTIs displace water molecules to do this, which provided them with more information to let them know what makes INSTIs clinically successful.
Lyumkis says, “In previous structures, we understood the biological properties of HIV intasome. But in this paper, we really begin to gain insight into how INSTI drugs target these important viral intasomes from a therapeutic perspective.”
These researchers are planning to carry out more studies on these experimental drugs, especially compound 4d. Based on preclinical testing and new structural insights, this compound is more expected to be resistant to HIV than other compounds. They also want to better understand what changes occur in the structure of HIV intasome in the event of resistance to INSTI drugs. Lyumkis says this may help them design more effective drugs in the future.
Reference
1.Dario Oliveira Passos et al. Structural basis for strand transfer inhibitor binding to HIV intasomes. Science, 2020, doi:10.1126/science.aay8015.
2.Imaging study of key viral structure shows how HIV drugs work at atomic level. https://www.salk.edu/news-release/imaging-study-of-key-viral-structure-shows-how-hiv-drugs-work-at-atomic-level/