In this study, researchers used the single LCNAN particle freezing electron microscopy to explore the mechanism. Since the quaternary complex of NALCN-FAM155-UNC79-UNC80 is not stable enough, they focus on the structural analysis of the relatively stable core subunit of NALCN-FAM155. After homologous protein screening and other steps, they determined the complex composed of rat NALCN and mouse FAM155a subunit as the research object. After overcoming the difficulties of sample preparation and data processing, the 2.65Å electron density of the NALCN-FAM155a complex was measured with high resolution, and the atomic model was built according to the homologous protein structure.
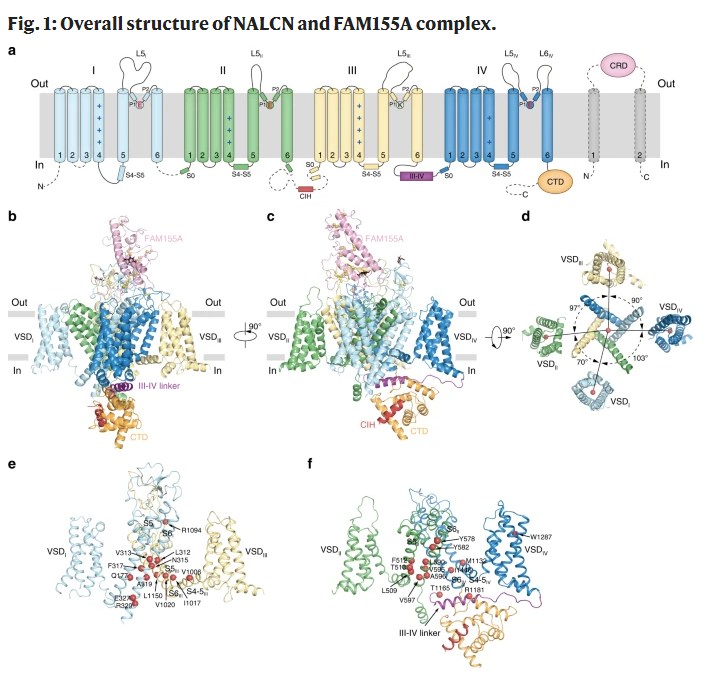
The overall structure of NALCN is similar to CAV and NAV, and consists of four homologous but different repeat domains (DI-DIV). Each repeat contains six transmembrane helices (S1-S6). The central S5-S6 forms the pore region, while the peripheral S1-S4 forms the piezoelectric sensing domain (VSD). The cysteine-rich domain (CRD) of extracellular FAM155a covers the top of the pore domain of NALCN like a lid and has complex and close interaction with the extracellular loop of NALCN. In addition, the C-terminal domain (CTD) of NALCN, III-IV linker, and CIH of DII-DIII form the intracellular domain.
By calculating the path of ions passing through, the researchers found that the amino acids around the extracellular pore region of NALCN have more negative charges, which are beneficial to attract extracellular sodium ions and repel anions. The calculated pore radius shows that the pore area shrinks tightly in two places: one is the “gate” close to the intracellular hydrophobic, indicating that the channel is closed; the other is the ion-selective filter close to the extracellular, which is responsible for selectively permeating sodium ions. Sequence alignment results showed that the ion-selective filter of NALCN was composed of “EEKE”, which was different from “DEKA” of NAV and “EEEE” of CaV1.1. High-resolution electron density shows that the selective filter amino acid E1389 of the fourth domain of NALCN has two conformations, but neither of them is located in the high field strength (HFS) layer of the selective filter, resulting in the similar charge distribution between NALCN and NAV in HFS layer, which may explain why NALCN has higher selectivity for sodium ions. In addition, some studies have shown that D1390 is more important than E1389 in blocking the current of NALCN by extracellular calcium ions, which is consistent with the observed structure.
The current of the NALCN channel complex is also regulated by voltage, in which VSD plays an important role. The function of VSD depends on its internal charge transfer center (CTC) and the continuous positively charged amino acids (R1-R6) on S4. CTC is mainly composed of one aromatic amino acid on S2, two negatively charged amino acids (E1-F3-E4), and the adjacent negatively charged amino acids on S3. Compared with voltage-gated ion channels such as NAV and CAV, VSD of NALCN degenerates to different degrees. According to the sequence alignment and structure comparison, it can be found that VSDI and VSDII of NALCN have complete CTC and positive charged amino acid cluster on S4, while CTC of VSDIII is not complete and lacks key aromatic amino acids on S2; although VSDIV has complete CTC, S4 has only two positive charged amino acids. Previous studies have shown that the positive amino acids on S4 of VSDI and VSDII are essential for the activity of NALCN, while the positive amino acids on S4 of VSDII and VSDIV are optional, which is consistent with the observed structure.
In conclusion, the high-resolution structure of the mammalian NALCN-FAM155a subcomplex was observed by structural biology techniques, which laid a foundation for further understanding of sodium selectivity, extracellular calcium blocking, and voltage regulation of NALCN.
Reference
- Yunlu Kang, Jing-Xiang Wu & Lei Chen. Structure of voltage-modulated sodium-selective NALCN-FAM155A channel complex. Nature Communications volume 11, Article number: 6199 (2020)