Saturation Transfer Differences (STD) NMR
Creative Biostructure provides MagHelix™ STD NMR technique for characterization of the binding epitope of a ligand to its receptor protein. STD NMR is a powerful and fast method used to study protein-ligand interactions, focusing on the signals of the ligand. Besides, values including KD, the association and dissociation kinetics (Koff and Kon respectively) can be obtained from STD NMR study as well.
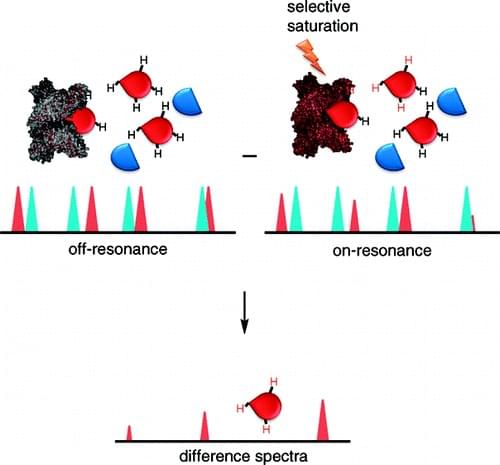
Figure 1: The scheme of the STD-NMR
In the context that in drug discovery, ligand-based NMR screening and the bound conformation of a ligand through NMR are important tools, now the technique of saturation transfer difference NMR (STD-NMR) has been widely used to characterize ligand-receptor complexes for many years. STD-NMR is a simple and fast method to screen ligand and characterize protein binding and it does not require expensive stable isotope or radioisotope labeling. Even to monitor low receptor concentration at low molecule concentration, STD can be used via non-scalar magnetization transfer, while NMR spectroscopy requires a much higher concentration of proteins that are labeled stable isotopes. In an STD-NMR experiment (as shown in the figure), the difference spectrum is obtained by subtracting a spectrum where the protein is selectively saturated (on-resonance) with signal intensities ISAT from that without saturation (off-resonance) with intensities I0. And the difference in intensities (ISTD=I0-ISAT) indicates molecule binding. There is possibility that other compounds may be present in the medium but only ligands can bind to the receptor and receive saturation transfer from the proteins to remain the signals. It is notoriously that membrane protein is very difficult to generate in exactly functional form. Due to the advantages of STD that it is sensitive enough to detect ligand binding at low protein concentration (pM–nM) similar as that in cells, there is no need to use bacterial expression systems to produce high protein yields. Moreover, membrane-bound proteins from cells without unfolding and refolding can be obtained under nondestructive conditions. Up to now, STD NMR has been used to monitor ligand binding to membrane proteins, such as members of G-protein-coupled receptors.
Our MagHelix™ analytical methods for biophysical characterization of biomolecules include but are not limited to:
- Protein Thermal Shift Assay
- Surface Plasmon Resonance
- Isothermal Titration Calorimetry
- Differential Scanning Calorimetry
- Saturation-Transfer Difference
- Bio-Layer Interferometry (BLI) Technology
AldinoViegas, et al. Saturation-transfer difference (STD) NMR: a simple and fast method for ligand screening and characterization of protein binding. Journal of Chemical Education, 2011, 88 (7): 990-994.
Venkitakrishnan RP, et al. Use of NMR Saturation Transfer Difference Spectroscopy to Study Ligand Binding to Membrane Proteins. Methods Mol Biol. 2012;914:47-63.
Ordering Process

