Negative Staining Electron Microscopy for Coronavirus-like Particle
High-resolution microscopy is essential for visualizing viral particles, and the acquisition of morphological characteristics of viral particles depends on the resolution. Transmission electron microscopy (TEM) is the starting point for obtaining the optimal image resolution. Negative staining electron microscopy (EM) of virus suspensions can provide detailed information on the architecture of viral particles. This is a technique that can be performed rapidly and can adapt to the highest magnification of viral particles. It also provides an initial model for structural analysis of cryo-electron microscopy (cryo-EM). Creative Biostructure is a leading supplier in the field of structural biology and offers you negative staining EM services for the visualization of coronavirus-like particles.
Application of Negative Staining EM in Virology
Because the main components of viruses are carbon-containing light element organics, such samples are easily passed through by high-energy electron beams, resulting in low contrast, which is not suitable for direct observation. Therefore, the morphological observation of viruses generally adopts the negative staining imaging technique, that is, it is necessary to impregnate the sample with heavy metal salts such as Pb, W, and U before the observation to achieve the structural enhancement for TEM imaging. Negative staining EM is a rapid and intuitive technique that is critical for determining the etiology of the large-scale outbreak of severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS). In addition, ultra-thin sections of virus-infected cell cultures or tissues incorporating a positive staining technique can provide information about the localization of the virus in the cellular environment. These two complementary techniques can be used to study the architecture of a virus and its parasitic life cycle.
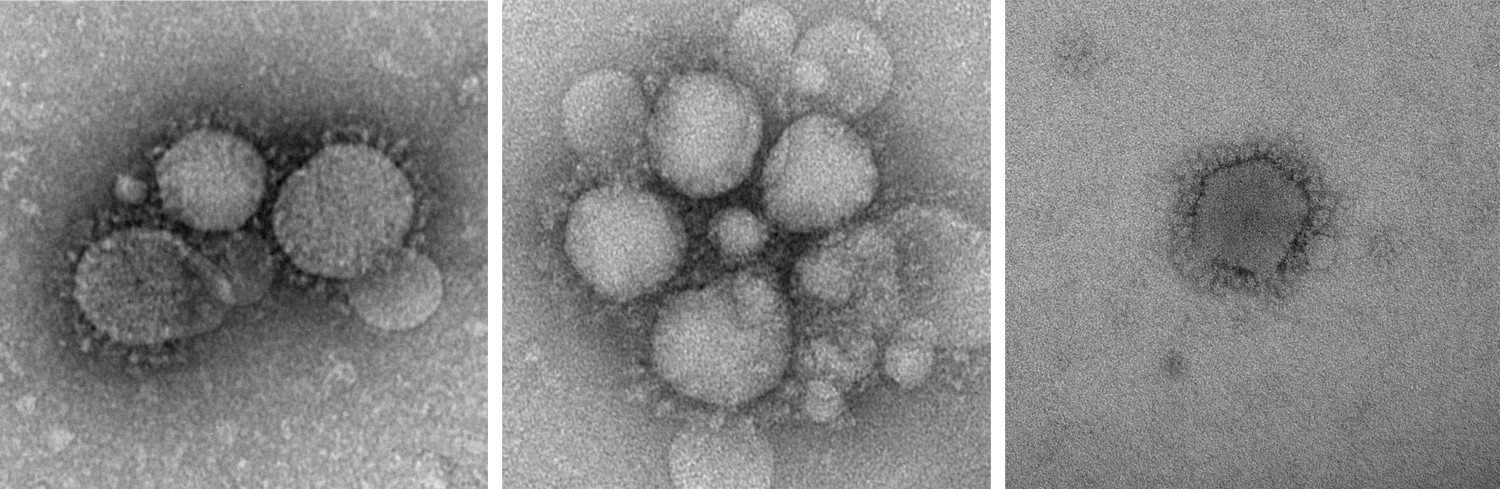 Figure 1. MERS-CoV particles as observed by negative staining EM. (CDC: Cynthia Goldsmith/Azaibi Tamin)
Figure 1. MERS-CoV particles as observed by negative staining EM. (CDC: Cynthia Goldsmith/Azaibi Tamin)
Our Negative Staining EM Services for Coronavirus-like Particle
Creative Biostructure does not offer diagnostic tests and the service is for research only. Moreover, for safety reasons, we do not accept some samples of coronavirus that could cause large-scale infection in humans. The general workflow of our negative staining EM is as follows:
- A drop of suspension of coronavirus/coronavirus-like particle is placed on a petri dish
- A Formvar-coated metal EM grid is placed Formvar side down on top of the drop of sample suspension for ~1-3 mins
- Remove the metal grid, blotted with filter paper and placed onto a drop of phosphotungstic acid (2.0%, pH 7.0) for one minute
- Blot up the excess phosphotungstic acid
- The metal grid adsorbed with sample suspension can be observed and photographed by TEM
Our negative staining EM service can be combined with the single-particle cryo-EM service, and our experts in three-dimensional (3D) structure reconstruction can use the image data of negative staining EM to build an initial model for single-particle analysis (SPA). In addition, more comprehensive information on viral structure and localization can be obtained by combining with ultra-thin section positive staining EM technology.
If you are interested in our negative staining EM services for coronavirus-like particles, please do not hesitate to contact us for more details. And our customer service representatives are available 24 hours a day from Monday to Sunday.
Contact us to discuss your project!
Reference
- Barreto-Vieira D F, Barth O M. Negative and positive staining in transmission electron microscopy for virus diagnosis. InTech, 2015.

 Figure 1. MERS-CoV particles as observed by negative staining EM. (CDC: Cynthia Goldsmith/Azaibi Tamin)
Figure 1. MERS-CoV particles as observed by negative staining EM. (CDC: Cynthia Goldsmith/Azaibi Tamin)