Protein Structure Analysis Using Negative Stain Electron Microscopy for Coronavirus Research
Negative stain electron microscopy (EM) is a technology that provides nanometer-resolution images of biological macromolecules. At present, the main role of negative stain EM in protein structural analysis is to construct an initial model and to conduct preliminary screening of samples before cryo-electron microscopy (Cryo-EM) observation. By applying the advances in cryo-EM image processing technology to the analysis of negative staining EM images, we can determine domain-level molecular structures (~10 Å) faster and more efficiently. Creative Biostructure currently supports the structural analysis of proteins and protein complexes involved in the process of coronavirus infection using negative staining EM.
Brief Introduction to Coronavirus
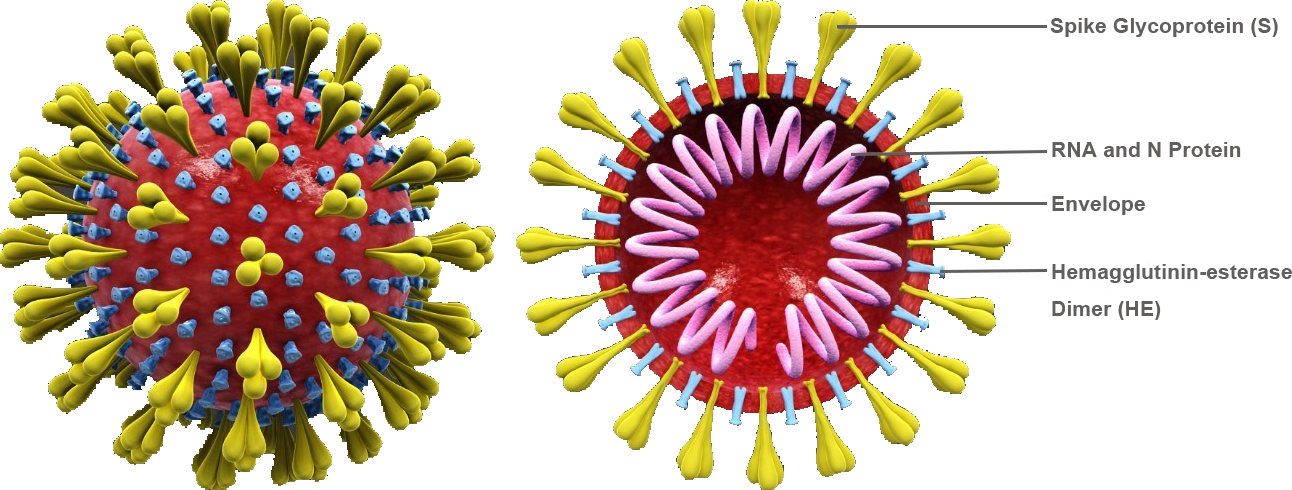
Coronaviruses (CoVs) are enveloped viruses that cause different diseases in humans and animals. Mouse hepatitis virus (MHV) causes hepatitis, enteritis and central nervous system diseases in rodents and is one of the most intensively studied CoVs. MHV belongs to the genus Betacoronavirus. Members from the same genus also include highly pathogenic CoVs, such as severe acute respiratory syndrome coronavirus (SARS-CoV), Middle East respiratory syndrome coronavirus (MERS-CoV) and novel coronavirus that have caused global pandemics. Coronavirus has a single-stranded positive RNA genome of approximately 30 kb, encoding 4-5 structural proteins, including nucleocapsid phosphoprotein (N), matrix (M), envelope protein (E), spike glycoprotein (S) and hemagglutinin-esterase (HE) of certain beta CoVs.
Why Do You Need This Service?
Due to heavy metal staining, the contrast of negative-stain image is high, and the preparation of negative-stain sample is simple. The information of macromolecular image analysis from negative stain EM is significant. First, information about the macromolecular morphology, organization, and heterogeneity of proteins or complexes can be obtained. Secondly, the negative-stain EM structural data of a larger number of samples can be obtained in a short time than that of Cryo-EM, so that the most promising samples can be screened by negative stain EM before using Cryo-EM to resolve atomic resolution structures. Finally, structural information from negative stain EM can be combined with other information such as antibody competition data, to in-depth understand the structure of macromolecules.
What We Can Do?
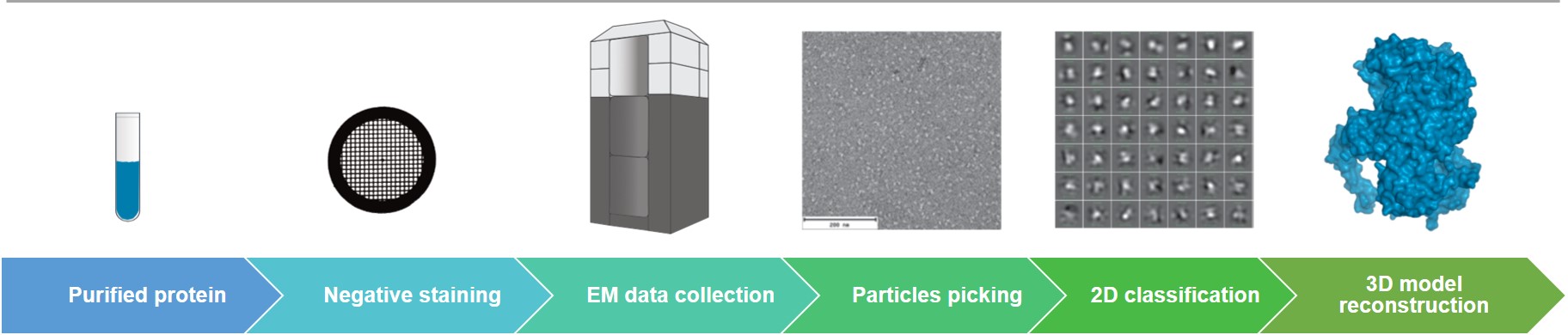
Creative Biostructure can efficiently perform sample preparation, data collection and data analysis of negative stain EM. The data processing software we use supports the automatic picking and classification of tens of thousands of particles. Our negative stain EM service provides the domain-level macromolecular structural analysis, such as epitope mapping, in studies of coronavirus infection.
Negative stain EM is a convenient method for assessing the quality of purified biomacromolecules at the microscopic scale. The purpose of the negative stain EM screening is to qualitatively evaluate the particle composition and conformational homogeneity. If you have chosen our single particle Cryo-EM service, we can provide you with negative stain EM service free of charge to screen out high-quality samples for Cryo-EM.

Why Do You Choose Us?
- State-of-the-art equipment
- Extensive knowledge and experience in electron microscopy
- Regular feedback and all-embracing service experience
- Flexible and high-quality service with competitive price
- Our customer service representatives are available 24 hours a day from Monday to Sunday
Contact us to discuss your project!
References
- Gui M.; et al. Electron microscopy studies of the coronavirus ribonucleoprotein complex. Protein & Cell. 2017, 8(3): 219-224.
- Gallagher J R.; et al. Negative-stain transmission electron microscopy of molecular complexes for image analysis by 2D class averaging. Current Protocols in Microbiology. 2019, 54(1): e90.



