Since December 2019, multiple cases of pneumonia with unknown etiology have been found in Wuhan, China for the first time. The World Health Organization (WHO) eventually terms the pneumonia as Coronavirus Disease 2019 (COVID-19), and the novel coronavirus causing the outbreak is named as Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) by International Committee on Taxonomy of Viruses (ICTV). COVID-19 has spread to all provinces in China and many other countries. As a leading supplier in the field of biopharmaceuticals, Creative Biostructure has established an advanced COVID-19 discovery platform with our expertise in structural biology, which can provide a wide range of contract research services involved in structure-based basic research, antiviral drug discovery, and vaccine development to help combat SARS-CoV-2 outbreak.
Taxonomy and Genome
Using next-generation sequencing (NGS) and metagenomic analysis, scientists have obtained several SARS-CoV-2 (provisionally named 2019-nCoV) full-length genome sequences, and their identity with each other is greater than 99.9%. The SARS-CoV-2 genome consists of six major open reading frames (ORFs) shared by coronaviruses and some other accessory genes, and amino acid sequence analysis of the seven conserved domains used for the classification of coronavirus species indicates that this virus may belong to the species of SARS-related coronavirus (SARSr-CoV).
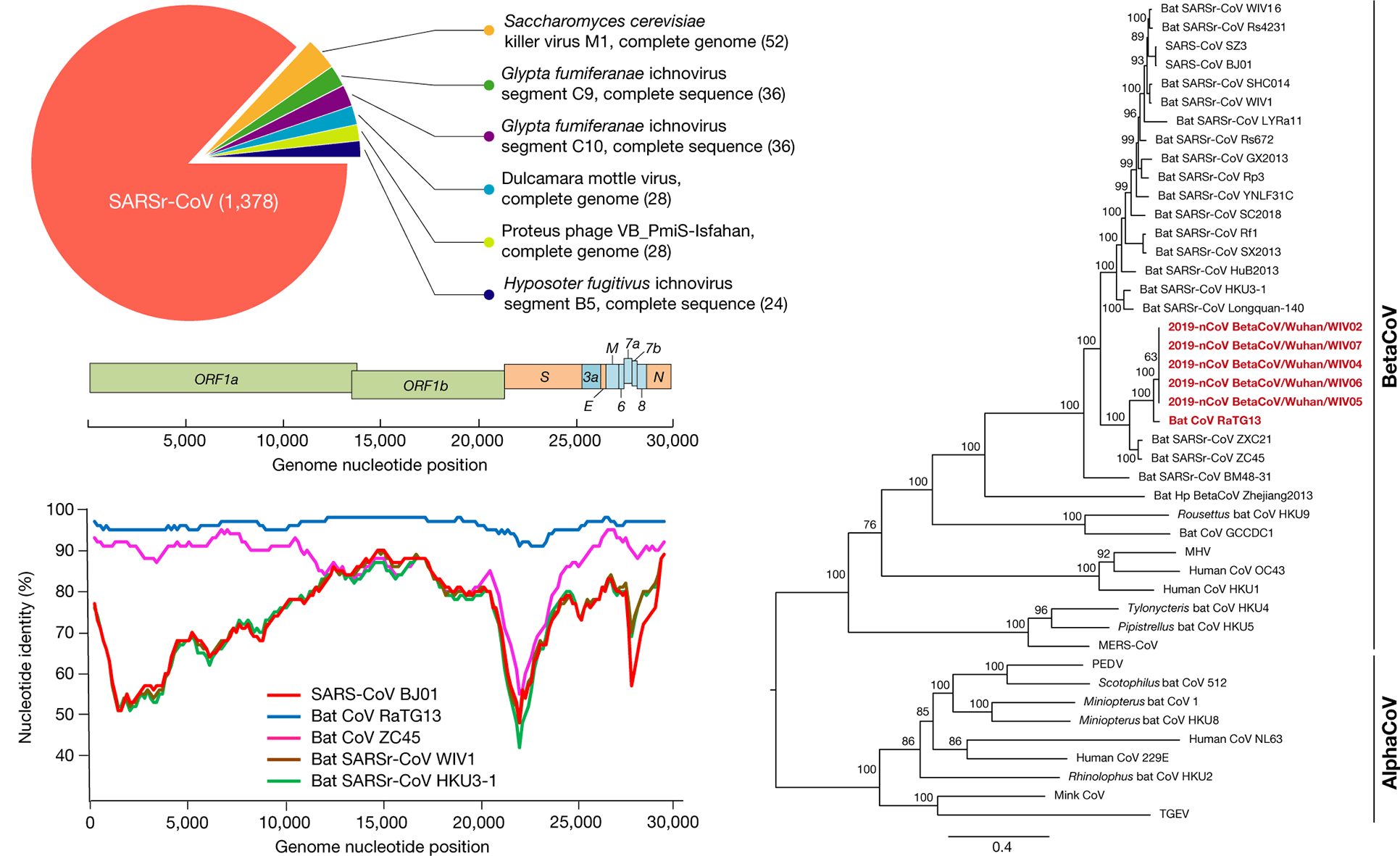 Figure 1. Genome characterization of SARS-CoV-2. (Zhou P.; et al. 2020)
Figure 1. Genome characterization of SARS-CoV-2. (Zhou P.; et al. 2020)
Structural Biology
SARS-CoV-2 is an enveloped, positive-sense, single-stranded RNA virus, and has similar structural characteristics to other coronaviruses. Coronavirus particles contain four major structural proteins, namely the nucleocapsid (N) protein, the spike (S) protein, the membrane (M) protein, and the envelope (E) protein. The main function of N protein is to protect and encapsulate the viral RNA forming viral RNP complex. The S protein forms a protruding structure on the surface of the virus. These protruding structures bind to receptors on the host cell, determining the types of cells the virus can infect and the range of species it can invade. Both the M protein and the E protein are thought to be involved in the assembly and release of the virus. The major nonstructural proteins of SARS-CoV-2 include 3-chymotrypsin-like protease (3CL), papain-like protease, helicase, and RNA-dependent RNA polymerase (RdRp).
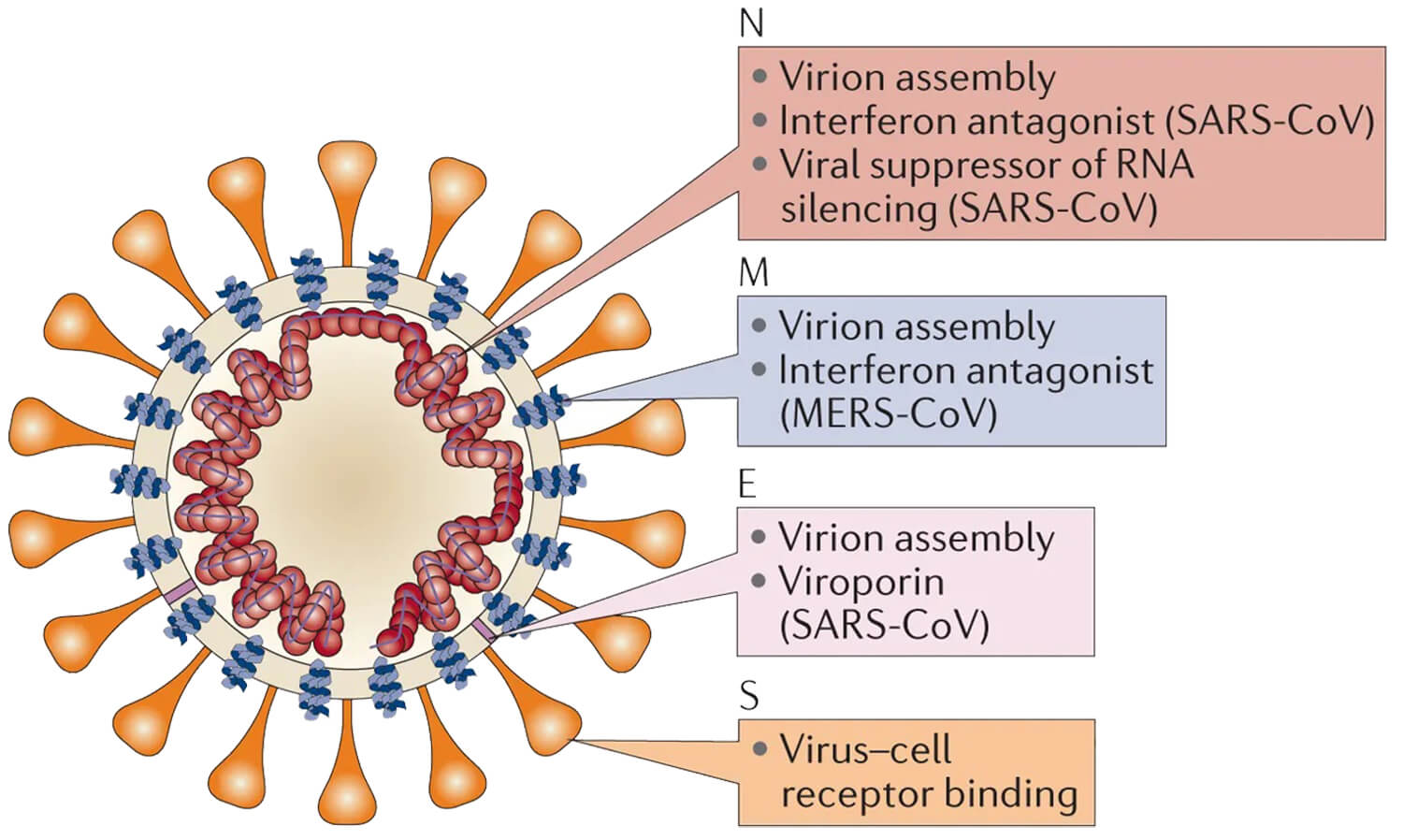 Figure 2. Structure of Coronavirus. (Adapted from Zumla A.; et al. 2016)
Figure 2. Structure of Coronavirus. (Adapted from Zumla A.; et al. 2016)
Using cryo-electron microscopy (cryo-EM) technology, scientists have successfully constructed the high-resolution structure (3.5 Å) of the S protein trimer on the surface of SARS-CoV-2, which will accelerate the development of antiviral drugs and vaccines. Moreover, the crystal structure of SARS-CoV-2 3CL hydrolase (Mpro) with high resolution has been obtained. This protease is crucial for proteolytic maturation of the virus and regulates the activity of the viral replication complex and is considered as a potential target for small-molecule drug development.
Discovery Platform for COVID-19 at Creative Biostructure
Creative Biostructure is committed to determine and analyze the structure of biomacromolecules. We have established a first-class discovery platform that targets SARS-CoV-2 to offer services (including but not limiting to) for structure/function study, vaccine development and novel drug discovery. Below is a list of our featured services:
 Figure 3. Our discovery platform for SARS-CoV-2/COVID-19.
Figure 3. Our discovery platform for SARS-CoV-2/COVID-19.
Contact us to discuss your project!
References
- Zhou P.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020: 1-4.
- Zumla A.; et al. Coronaviruses-drug discovery and therapeutic options. Nature Reviews Drug Discovery. 2016, 15(5): 327.

 Figure 1. Genome characterization of SARS-CoV-2. (Zhou P.; et al. 2020)
Figure 1. Genome characterization of SARS-CoV-2. (Zhou P.; et al. 2020) Figure 2. Structure of Coronavirus. (Adapted from Zumla A.; et al. 2016)
Figure 2. Structure of Coronavirus. (Adapted from Zumla A.; et al. 2016) Figure 3. Our discovery platform for SARS-CoV-2/COVID-19.
Figure 3. Our discovery platform for SARS-CoV-2/COVID-19.