Creative Biostructure has established an excellent platform after years of membrane protein production experience. Our custom Mempro™ T-cell surface glycoprotein CD3 zeta chain production service is based on our cell-based expression system.
Creative Biostructure can apply the most advanced method-Mempro™ cell-based protein production system to perform T-cell surface glycoprotein CD3 zeta chain. CD3 zeta chain is one of the glycoproteins expressed on T cell plasma membrane, also as known as CD247. CD3 zeta, CD3 gamma, delta, and epsilon chain, with CD3 alpha/beta and CD3 gamma/delta heterodimers, can together form the T cell receptor CD3 complex. CD3 zeta chain plays a crucial role in recognizing the coupling antigen in several signal transduction pathways through its ITAM signaling module. Low expression of CD3 zeta chain coupling antigen leads to impaired immune response. Lipid-protein interaction can apparently regulate ITAM to switch between functional and non-functional conformations. Unlike CD3 epsilon and CD3 gamma, CD3 zeta chain has not present polybasic regions next to their ITAM motifs, therefore its cytoplasmic tail cannot interact with the plasma membrane.
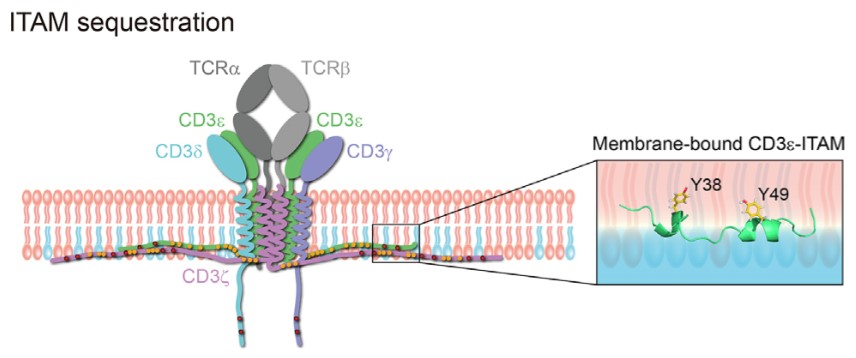 Figure 1. Self-assembly for TCR CD3 and direct consequence of CD3-lipid interaction. (Progress in Biophysics and Molecular Biology, 2015)
Figure 1. Self-assembly for TCR CD3 and direct consequence of CD3-lipid interaction. (Progress in Biophysics and Molecular Biology, 2015)
Creative Biostructure is expertized in membrane protein production services. Mempro™ cell-based protein production platform provide multiple choices to accomplish your goal in different systems, including:
• Mempro™ CD3 zeta chain Production in Mammalian Cells System
Creative Biostructure can provide another great service-Mempro™ protein production in mammalian cells system. This system helps to correct folding and post-translational modifications for membrane protein. Creative Biostructure provides different approaches to produce high-quality membrane protein in mammalian cells:
1. Easy and rapid production-Transient transfection system;
2. Higher yield and reproducible production-Stable transfection system;
3. Adenovirus-assist protein expression.
• Mempro™ CD3 zeta chain Production in Bacterial Cells System
Escherichia coli (E. coli) is the most widely employed bacterial host for membrane protein production. Lemo21 (DE3) strain is optimized for large amount Mempro™ protein production.
• Mempro™ CD3 zeta chain Production in Insect Cells System
Creative Biostructure offers our advanced Mempro™ membrane protein production using insect cells system. This system is widely used for membrane protein production in cultured baculovirus system. According to similar codon usage rules, Mempro™ protein production using insect cells system can increase the expression level and reduce the truncated proteins compared to E. coli system.
• Mempro™ CD3 zeta chain Production in Yeast Cells System
Single-celled yeast is an easy, quick and economic culture host and able to apply multiple post translation modifications for eukaryotic membrane protein. Creative Biostructure has developed several strategies to improve yields per cell through optimizing the expression plasmid, host cell and culture conditions.
Creative Biostructure provides Mempro™ cell-based protein production services including expressing, isolating, purifying and crystallizing the T-cell surface glycoprotein CD3 zeta chain to facilitate its further studies on T cell signaling mechanism.
We provide an array of Mempro™ membrane protein production services. Please feel free to contact us for a detailed quote.
References:
W. Wei, et al. (2015). Lipid in T-cell receptor transmembrane signaling. Progress in Biophys. and Mol. Biol, 118(3): 130-138.
A. B. Sigalov, et al. (2006). Lipid-binding activity of intrinsically unstructured cytoplasmic domains of multichain immune recognition receptor signaling subunits. Biochemistry, 45: 15731-15739.
L. Li, et al. (2014). Ionic protein-lipid interaction at the plasma membrane: what can the charge do? Trends Biochem. Sci., 39: 130-140.