Exosome Isolation Services
Exosomes are tiny vesicles secreted by most cells with a lipid bilayer structure, found in various body fluids like blood, urine, saliva, and more. They carry significant information, such as proteins, lipids, and non-coding RNAs (miRNA, lncRNA, circRNA, etc.), making them crucial for cell-to-cell communication and potential early diagnostic markers for diseases. Exosome RNA sequencing can quickly and efficiently provide comprehensive information about exosome RNA, making it an ideal tool for disease diagnosis and follow-up, focusing on miRNA, lncRNA, circRNA, mRNA, and other RNA types.
Exosomes are present in biological and cell culture fluids but often mixed with cell debris and protein aggregates. Compared to nucleic acids, detecting differential expression of exosome proteins is challenging due to contamination risks with conventional isolation methods. Creative Biostructure can isolate and purify high-purity exosomes for biological analysis.
Exosome Isolation Services from Various Sources

- Human and animal tissues, such as adipose tissue, brain tissue, liver tissue, kidney tissue, muscle tissue, bone tissue, and connective tissue.
- Plant tissues, such as leaves, roots, stems, seeds, fruits, and flowers.
Our Methods for Exosome Isolation and Purification
Creative Biostructure provides various methods for exosome isolation, including traditional differential centrifugation, density gradient centrifugation, and more recent methods such as ultrafiltration, polymer precipitation, immune separation, isolation screening, and size exclusion chromatography.
Differential Ultracentrifugation (dUC)
| Diagram | Principle | Technical Features |
|
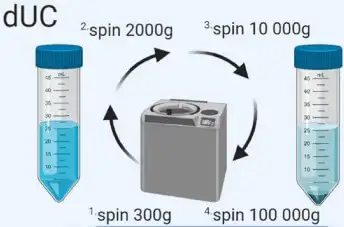
It is a common exosome separation technique involving several steps: - Low-speed centrifugation to remove cell and apoptotic fragments. - Higher-speed centrifugation to eliminate larger vesicles. - High-speed centrifugation to precipitate exosomes. |
Exosomes can be extracted from cell culture supernatant, serum, plasma, urine, cerebrospinal fluid, and other body fluids (ascites, amniotic fluid, emulsion, saliva, etc.), and the obtained high-purity exosome granules can be used for subsequent NTA, TEM, WB analysis, etc. However, there is a risk of co-precipitating non-exosomal materials and potential damage to exosomes due to high forces. |
Size-Exclusion Chromatography (SEC)
| Diagram | Principle | Technical Features |
|
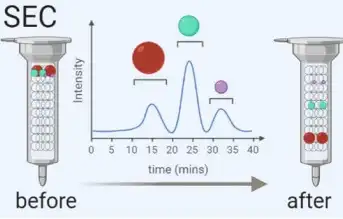
SEC separates macromolecules based on size using columns filled with porous polymer microspheres. Smaller molecules take longer to pass through the pores, while larger molecules elute earlier. | SEC has advantages over centrifugation as it avoids shear forces on exosomes. It is widely used for isolating exosomes from blood and urine. We offer standard chromatography-based procedures to ensure high recovery rates of exosomes. |
Poly-Ethylene Glycol (PEG)-Based Precipitation
| Diagram | Principle | Technical Features |

|
Exosome precipitation from biological fluids is achieved by altering solubility using a PEG-containing solution. After overnight incubation, exosomes are isolated by low-speed centrifugation or filtration. | This method is easy, requires no specialized equipment, and is scalable for many samples. Various precipitation kits are available. However, there is a risk of polymer contamination and potential disruption of exosome integrity. |
Density Gradient Centrifugation (DGC)
| Diagram | Principle | Technical Features |
|
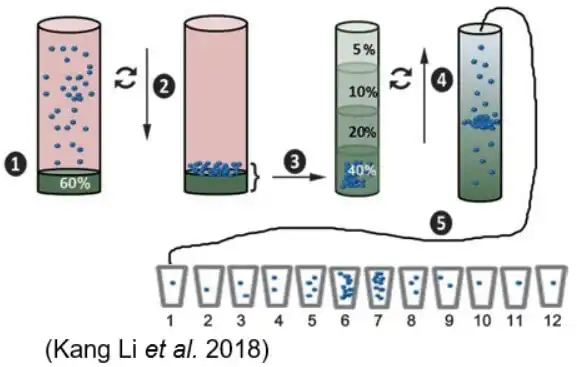
Density gradient centrifugation uses ultracentrifugation with a sucrose gradient to separate exosomes from non-vesicular particles, such as proteins and protein/RNA aggregates. | It effectively isolates exosomes by density but requires precise centrifugation times to avoid contamination by particles of similar density. |
Immunoaffinity (IA)
| Diagram | Principle | Technical Features |
|
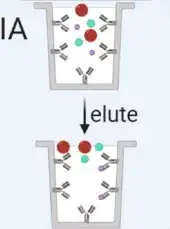
Immunoaffinity capture using beads coated with antibodies involves several experimental procedures, including sample treatment, bead incubation and binding, low-speed centrifugation, and biological analysis of the isolated exosomes. | Using beads coated with antibodies that target exosome surface proteins allows for the isolation of specific vesicle populations. However, this method is limited by the availability of well-characterized antibodies and may result in the loss of non-targeted exosomes. |
Microfluidics (MF)
| Diagram | Principle | Technical Features |
|
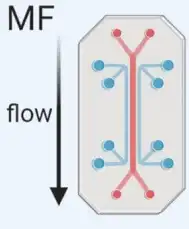
Microfluidics (MF) technology uses chips with specific antibody-mediated binding to capture exosomes. With a microfluidics-based platform, we can provide high throughput isolation and purification services for our customers. | The Microfluidics-based technique can be used in exosome isolation with specific surface markers (microfluidics combined with immunoaffinity capture) and in exosome isolation with specific size populations (microfluidics combined with membrane filtration). |
Ultrafiltration (UF)
| Diagram | Principle | Technical Features |
|
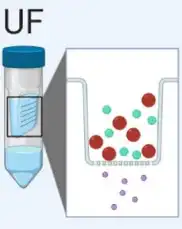
Ultrafiltration (UF) is dependent on a filter with different pore sizes that are used to separate exosomes from proteins and other macromolecules. | Simple and quick, this method is suitable for large-volume samples. However, it may co-isolation with contaminants. |
Active Exosome Isolation
Our active exosome isolation method addresses the limitations of traditional gradient centrifugation, which often compromises exosome morphology and biological activity. Our optimized technique uses a special material to separate and purify exosomes from samples, ensuring they maintain their complete morphology and biological activity. This rapid and cost-effective method yields morphologically intact exosomes that are crucial for drug delivery, exosome research, and immunotherapy, demonstrating excellent application potential.
Technology Platforms
Key Advantages
- Support Multiple Sources: We offer extraction from various biological sources, including animal-derived, plant-derived, and microorganism-derived exosome isolation.
- High Purity: Our methods yield high-purity exosomes suitable for downstream applications such as NTA, TEM, and WB analysis.
- Technique Diversity: We employ multiple techniques such as differential centrifugation, density gradient centrifugation, and precipitation methods to cater to different sample types and research needs.
- Scalability: Our methods are scalable for processing both small and large sample volumes, ensuring flexibility in experimental design.
- Specialized Equipment: We utilize specialized equipment to minimize contamination and ensure the integrity of isolated exosomes.
- Expertise: Our team has extensive experience in exosome research and purification techniques, ensuring reliable and reproducible results. We can offer customizable protocols tailored to specific research requirements.
Resources
Frequently Asked Questions
-
How do you ensure the purity and integrity of isolated exosomes?
We use advanced techniques such as differential centrifugation, density gradient centrifugation, and precipitation methods to ensure high-purity exosome preparations. Our specialized equipment minimizes contamination, and our protocols are designed to preserve the integrity of the exosomes, avoiding potential damage from high forces.
-
Can you customize the exosome isolation protocols to meet specific research needs?
Yes, we offer customizable protocols tailored to your specific research requirements. Our team has extensive experience and can adjust the extraction and purification methods to suit various sample types and research goals, ensuring you receive the most relevant and reliable results.
-
How scalable are your exosome isolation services for large-volume samples?
Our exosome isolation methods are highly scalable, accommodating both small and large sample volumes. This flexibility ensures that we can handle varying experimental needs, from small-scale pilot studies to large-scale research projects, providing consistent and high-quality results regardless of sample size.
Ordering Process
References
- Karim Sidhom, et al. A review of exosomal isolation methods: is size exclusion chromatography the best option? Int. J. Mol. Sci. 2020, 21, 6466.
- Yong Kyoung Yoo, et al. Toward exosome-based neuronal diagnostic devices. Micromachines (Basel). 2018, 9(12): 634.
- Kang Li, et al. Cushioned-density gradient ultracentrifugation (C-DGUC): a refined and high performance method for the isolation, characterization, and use of exosomes. Methods Mol Biol. 2018, 1740: 69-83.
- Shang-Chun Guo et al. Microfluidics-based on-a-chip systems for isolating and analysing extracellular vesicles. J Extracell Vesicles. 2018, 7(1): 1508271.