Creative Biostructure has established the comprehensive membrane protein production platform based on years of experience, our scientists can provide custom Mempro™ transmembrane protein TYROBP production services using cell-based expression system.
Mempro™ cell-based protein production system is the one of our most applied and advanced method for membrane proteins production. TYROBP, also called DAP12, is well known for its crucial role of activating signal transduction in the natural killer cells. TYROBP consists of three parts: a short extracellular tail, a single transmembrane domain and a cytoplasmic domain with two signaling module ITAMs. Loss of function mutations in DAP12 gene were identified as the cause of a rare disorder Nasu-Hakola disease. TYROBP forms homodimer through polar interaction, and associates with an array of receptors to form a heterotrimeric activating immunoreceptor complex. NK cell activity was regulated by these complexes, which not only ensure protection of the hosts against pathogens, but also prevent the deleterious NK cell-driven autoimmune responses.
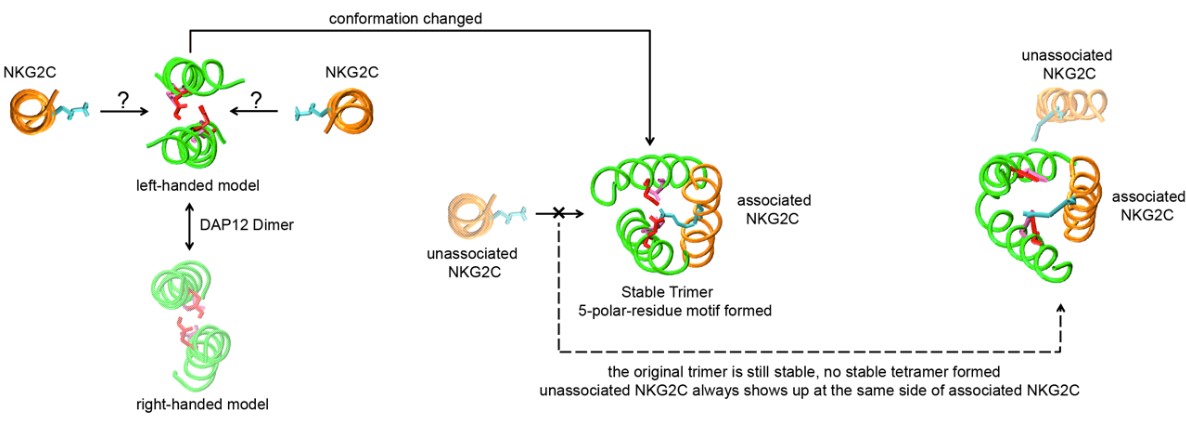 Figure 1. Self-assembly principle of actvating immunoreceptor TYROBP-NKG2C
Figure 1. Self-assembly principle of actvating immunoreceptor TYROBP-NKG2C
(Plos One, 2014)
Creative Biostructure provides high quality transmembrane protein TYROBP production, we can adopt various systems for Mempro™ cell-based protein production, including:
• Mempro™ TYROBP Production using Yeast Cells System
Single-celled yeast is quick, easy and economic to culture and capable to apply post-translational modification for eukaryotic membrane protein. With this popular membrane production alternative, Creative Biostructure can provide Mempro™ cell-based TYROBP production in yeast cells system.
• Mempro™ TYROBP Production using Incest Cells System
The baculoviruses are also widely applied for eukaryotic membrane protein production
in cultured insect cells and insect larvae. Creative Biostructure can provide Mempro™ cell-based TYROBP production in bacterial cells systems.
• Mempro™ TYROBP Production using Bacterial Cells System
For the production of membrane proteins, Escherichia coli (E. coli) is another widely applied bacterial host. We have optimized Lemo21 (DE3) strain for our Mempro™ protein production in E. coli system. Creative Biostructure can employ other host systems including Lactococcus lactis (L. lactis), Rodhobacter sphaeroides (R. sphaeroides), and Bacillus subtilis (B. subtilis) to producte TYROBP in bacterial cells systems.
• Mempro™ TYROBP Production using Mammalian Cells System
In order to perform correct membrane protein folding and post-translational modification, another great service Creative Biostructure can provide is Mempro™ protein production in mammalian cells system. We have proprietary Mempro™ protein production in mammalian cells system with high quality eukaryotic membrane proteins from numerous mammalian cell types including human embryonic kidney cells (HEK-293), Chinese hamster ovary cells (CHO), monkey kidney fibroblast cells (COS-7) and baby hamster kidney cells (BHK-21).
With the Mempro™ cell-based protein production platform, Creative Biostructure provides services including expressing, isolating, purifying and crystallizing TYROBP to accelerate the study of its function in multiple signal transduction pathways.
We provide an array of Mempro™ membrane protein production services. Please feel free to contact us for a detailed quote.
References:
P. Wei (2014). Molecular Dynamic Simulation of the Self-Assembly of DAP12-NKG2C Activating Immunoreceptor Complex. Plos One, 9(8): e105560.
J. C. Thrash, et al. (2008) Developmental Regulation of TREM2 and DAP12 Expression in the Murine CNS: Implications for Nasu-Hakola Disease. Neurochem Res., 34 (1): 38-45.
M. E. Call, et al. (2010). The structural basis for intramembrane assembly of an activating immunoreceptor complex. Nature Immunology, 11(11): 1023-1029.
A. S. Tessarz, and A. Cerwenka. (2013). The TREM-1/DAP12 pathway. Immunology Letters, 116(2): 111-116.
