Creative Biostructure provides the most advanced Mempro™ custom ion channel superfamily production services using virus-like particles (VLPs) for both industry and academic applications.
Ion channel proteins are important pore-forming membrane proteins that can be identified from membranes of nearly all cells. They can gate the flow of certain ions across the plasma membrane to establish a resting membrane potential, shaping action potentials or other electrical signals, regulating cell volume, and control the ion flows across the secretory and epithelial cells. Ion channel proteins differ from other types of ion transporters mainly in two features. First, ion channel proteins can achieve a significantly higher ion flow rate (106 per second or even higher). Second, the special ions that flow pass the membrane through ion channel proteins can only flow down their electrochemical gradient, and with no need for metabolic energy.
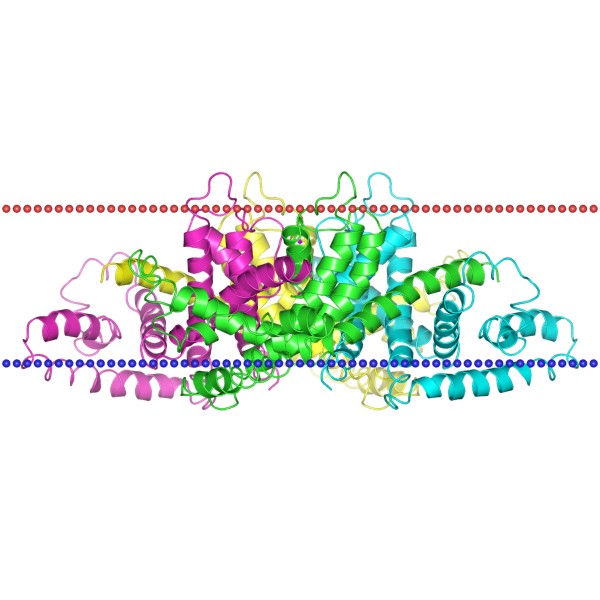 Figure 1. The schematic structure of Potassium channel KvAP (OPM Database)
Figure 1. The schematic structure of Potassium channel KvAP (OPM Database)
With the most advanced VLPs platform and years of experience in the field of recombinant membrane protein production, Creative Biostructure now can provide our state-of-art custom Mempro™ ion channel superfamily protein production service for various applications. Creative Biostructure have successfully provided recombinant protein production services in hundreds of cases with high quality these years. Our science team will develop unique and optimal strategy in each individual case according to the characteristics of target ion channel superfamily proteins and your special experimental requirements.
VLPs are highly organized subunits that can be described as "empty viral protein shells". They are constructed by the self-assembly of viral structural proteins. VLPs can mimic the native virus in many ways. But all the VLPs are non-infectious because of lacking the viral genetic materials. Many virus families have been used to obtain VLPs, such as Parvoviridae, bacteriophages, Retroviridae, etc. VLPs have been involved in various researches as powerful tools, including: vaccine development, heterologous epitope presentation, antigen delivery and recombinant membrane protein production.
VLPs are powerful tools for the production of recombinant membrane proteins. In a VLPs assay, we will co-express the target ion channel superfamily protein along with a certain viral structural core protein within the chosen host cells (bacteria, yeasts, green plants, insect cells and mammalian cells). The recombinant viral structural core protein will self-assemble within the plasma membrane of the host cells, and then bud-off from the plasma membrane to form the lipoparticles (VLPs that that contain high concentration of specific membrane proteins within their construction). During the formation of lipoparticles, the expressed target ion channel superfamily proteins will be captured within the construction of lipoparticles. The specific lipoparticles can be harvested at the surface of host cells with the target ion channel superfamily proteins being protected within its construction. The entire process is free from detergents and mechanical disruptions.
Creative Biostructure can also provide various other Mempro™ recombinant protein production services using multiple advanced methods. Welcome to contact us for more details.
Reference:
Sakamuro, D., Elliott, K. J., Wechsler-Reya, R., & Prendergast, G. C. (1996). BIN1 is a novel MYC–interacting protein with features of a tumour suppressor. Nature genetics, 14(1), 69-77.
Negorev, D., Riethman, H., Wechsler-Reya, R., Sakamuro, D., Prendergast, G. C., & Simon, D. (1996). The Bin1 Gene Localizes to Human Chromosome 2q14 by PCR Analysis of Somatic Cell Hybrids and Fluorescencein SituHybridization. Genomics, 33(2), 329-331.
A.Roldão, et al. Virus-like particles in vaccine development.Expert Rev. Vaccines, 2010,9(10): 1149-1176.
Feussner I, Wasternack C. The lipoxygenase pathway [J]. Annual review of plant biology, 2002, 53(1): 275-297.
