Creative Biostructure has established a first-class membrane protein production platform. Our customized Mempro™ MAPEG superfamily production service is based on our advanced cell-based expression systems, which express the target membrane protein production within a living cell.
MAPEG superfamily, short for membrane-associated proteins in eicosanoid and glutathione metabolism, is a type of integral membrane enzymes which enable to catalyze glutathione-dependent transformations of lipophilic substrates. Members of human MAPEG superfamily, including ALOX5AP, LTC4S, MGST1, MGST2, MGST3 and PTGES, conduct highly divergent functions. Recnet sequence studies indicated that there are four transmembrane helices serve as a common denominator of the MAPEG superfamily. In transmembrane helix 2, a highly conserved RX3NX2E/D motif is revealed in this significant conserved sequence. Recent structural studies showed that glutathione binds the LTC4S deep in its active site pocket in a horseshoe-shaped conformation, while the carboxylate moieties in glutathione is facing into the protein pocket, thus, exposing the thiol group to the membrane. The binding site for a horseshoe-shaped glutathione is assumed to be conserved throughout the MAPEG superfamily and the structural basis for the glutathione activation is considered to be alike among members of MAPEG superfamily as well. More structural information is still needed to shed light on the functions of MAPEG superfamily.
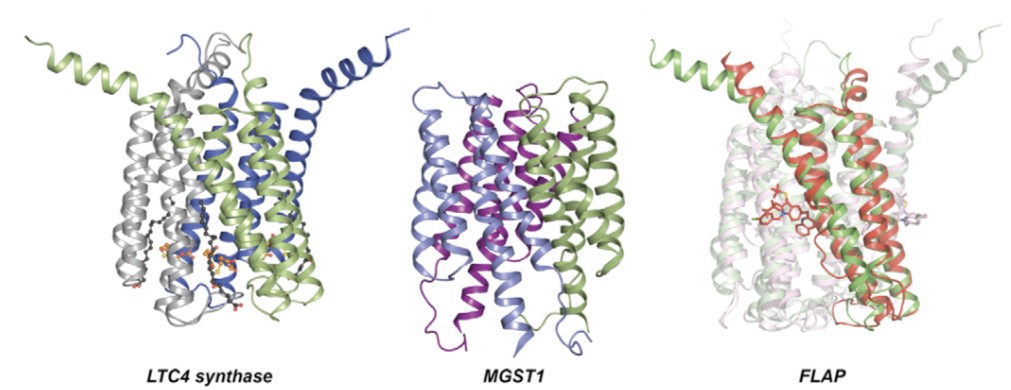
Figure 1.Currently available structures of the MAPEG superfamily (Curr. Opin. Struc. Biol., 2008)
Based on technology innovation, our cell-based expression platform is aimed to overcome the disadvantages in membrane protein production . Creative Biostructure can provide you the first-class membrane protein production services, and we can perform various Mempro™ cell-based protein production systems including:
• Mempro™ MAPEG Superfamily Production using Mammalian Cells System
Creative Biostructure can provide Mempro™ protein production in mammalian cells system. This system facilitates membrane proteins to perform correctly folding and post-translational modification. Various applications for eukaryotic membrane protein production by Mempro™ mammalian cells system lead to the best structural and functional features.
• Mempro™ MAPEG Superfamily Production using Insect Cells System
The most applied system for eukaryotic membrane protein production is in cultured insect cells and larvae. Mempro™ protein production using insect cells system can increase the expression level and reduce the truncated proteins compared to E. coli system according to similar codon usage rules. Our innovational insect cells system based on baculovirus is advanced in several aspects as below:
• Accuracy;
• Easy scale-up;
• High membrane protein yield;
• Proper folding and post translation modifications;
• Use of cell lines ideal for suspension culture.
• Mempro™ MAPEG Superfamily Production using Bacterial Cells System
Escherichia coli (E. coli) is the most widely applied bacterial host for membrane protein production. Lemo21 (DE3) strain is optimized for our Mempro™ protein production in our bacterial cells system. The target membrane protein encoding gene is located on a pET vector governed by the T7lac promoter. Membrane proteins are expressed as C-terminal GFP fusion protein as a result.
• Mempro™ MAPEG Superfamily Production in Yeast Cells System
Single-celled yeast is an easy, quick and economic culture host and able to apply multiple post translation modifications for eukaryotic membrane protein. Creative Biostructure has developed several strategies to improve yields per cell through optimizing the expression plasmid, host cell and culture conditions.
Creative Biostructure conducts high-yield MAPEG superfamily production by Mempro™ cell-based protein production services including expression, isolation, purification and crystallization.
We provide an array of Mempro™ membrane protein production services. Please feel free to contact us for a detailed quote.
References:
D. M. Molina, et al. (2008). Catalysis within the lipid bilayer-structure and mechanism of the MAPEG family of integral membrane proteins. Curr. Opin. Struc. Biol., 18: 442-449.
H. Hebert and C. Jegerschold. (2007). The structure of membrane associated proteins in eicosanoid and glutathione metabolism as determined by electron crystallography. Curr Opin Struct Biol, 17:396-404.
P. J. Jakobsson, et al. (2000). Membrane-associated proteins in eicosanoid and glutathione metabolism (MAPEG). A widespread protein superfamily. Am. J. Respir. Crit. Care Med., 161:20S-24.
