Creative Biostructure can provide custom Mempro™ P-loop containing nucleoside triphosphate (NTP) hydrolase production services using cell-based expression system.
Mempro™ cell-based protein production system is the most widely used for the production of membrane proteins.P-loop containing NTP hydrolase superfamily, which contains common sequence patterns (two distinct motifs), the Walker A motif (the P-loop proper) and Walker B motif that binds, respectively, the beta and gamma phosphate moieties of the bound NTP, and a Mg2+ cation. The P-loop containing NTP hydrolase fold is the most common domain of the several distinct nucleotide-binding protein folds. It is shown that P-loop NTP hydrolase shows substantial substrate preference for either ATP or GTP.
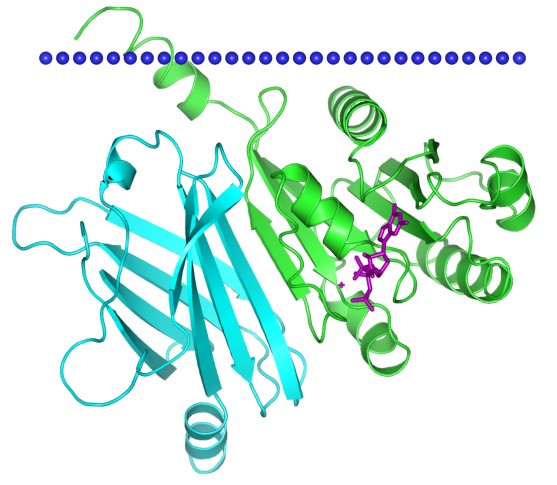
Figure 1. ADP-ribosylation factor-like protein 2, complex with human PDEdelta. (OPM database)
Creative Biostructure can produce high-yield P-loop containing NTP hydrolases, we can perform various strategies for Mempro™ cell-based protein production, including:
- Mempro™ Protein Production in Bacterial Cells System ;
- Mempro™ Protein Production in Yeast Cells System;
- Mempro™ Protein Production in Incest Cells System;
- Mempro™ Protein Production in Mammalian Cells System.
Escherichia coli (E. coli) is the powerful bacterial host for the production of P-loop containing NTP hydrolases. Yeast cells (such as saccharomyces cerevisiae and pichia pastoris) system, combines prokaryotic as well as eukaryotic characteristics. It is a perfect eukaryotic host due to single cells, fast growth rates, inexpensive media, high cell densities. Insect cells (such as Sf9, Sf21 and High Five) are also the major system for P-loop containing NTP hydrolase expression, which are developed from transfected insect cells in combination with vectors derived from the baculovirus species AcMNPV. Mammalian cells present another eukaryotic system for P-loop containing NTP hydrolase expression, which provide near-native cell environment. Cell lines derived from COS, CHO, BHK-21, HEK293, HeLa and GH3 can be generally used for membrane protein expression.
With the Mempro™ cell-based protein production platform, Creative Biostructure is capable of expressing, isolating, purifying and crystallizing P-loop containing NTP hydrolases to facilitate the study of their physiological functions.
We provide other various Mempro™ membrane protein production services. Please feel free to contact us for a detailed quote.
References:
F. Junge, et al. (2008) Large-scale production of functional membrane proteins. Cellular Mol. Life Sci., 65 (11): 1729-1755.
J. Petschnigg, et al. (2011). Using yeast as a model to study membrane proteins. Curr. Opin. Nephrol. Hypertens., 20(4): 425-432.
J. Sang, et al. (2006). Duplication and combination of P-loop containing nucleotide triphosphate hydrolases superfamily. Wuhan University J. Nat. Sci., 11(3): 577-580.
S. Schlegel, et al. (2013). Bacterial-based membrane protein production. Biochim. Biophys. Acta., 1843(8): 1739-1749.
Y. Kawamura, et al. (2003). Systematic analyses of P-loop containing nucleotide triphosphate hydrolase superfamily based on sequence, structure and function. Genome Informatics, 14: 581-582.