Directed Evolution
Directed evolution is one of the most popular strategies used for protein engineering. Creative Biostructure can perform directed evolution in vivo/vitro to evolve proteins with desired properties for our customers. Compared with rational design, it is not necessary to understand the protein structure in depth. Directed evolution mimics the natural evolution and harness the power of mutation and natural selection. After rounds of iterative enhancement, the protein(s) with desired functions will be produced.
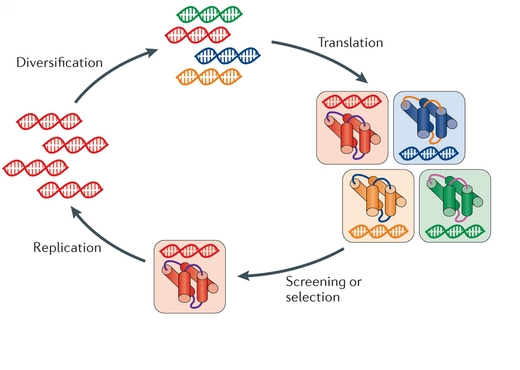 Figure: Key steps in the cycle of directed evolution. (Nat Rev Genet. 2015)
Figure: Key steps in the cycle of directed evolution. (Nat Rev Genet. 2015)
The method of directed evolution provides a choice to enhance protein performance under certain conditions. As shown in the figure and flow chart. Directed evolution for protein engineering is an iterative strategy. To determine a target biomolecule is the first step to start the procedure. Directed evolution mimics the natural evolution, so there are various in vitro strategies to generate diversity. The mainly to introduce diversity methods include in vitro mutagenesis (mutagenic strains, error-prone PCR, saturation mutagenesis, mutagenesis by random insertion or deletion), homologous recombination (DNA shuffling, family shuffling, oligonucleotide- and oligonucleotide primer-based methods, random chimeragenesis on transient templates (RACHITT), nucleotide exchange and excision technology (NExT), heritable recombination, genome shuffling) and non-homologous recombination methods (exon shuffling, incremental truncation methods, sequence Homology-independent protein recombination (SHIPREC), degenerate homoduplex recombination (DHR), sequence independent site-directed chimeragenesis (SISDC), random multi-recombinant PCR (RM-PCR) and Y-ligation based shuffling (YLBS)). After the diverse library of mutants is generated, high-throughput screening and selection methods are utilized to identify compounds with improved properties. Fluorescence-activated cell sorting (FACS) may be the most widely used high-throughput method for protein evolution, which identifies and isolates cells containing desired gene variants automatically but does not rely on diffusing fluorescent reporter. FACS can also be used combined with yeast display facilitating FACS to screen protein-protein interactions, bond formation and peptide bond cleavage. And the selection schemes include Phage, mRNA, ribosome and cell surface display, organismal fitness and auxotroph complementation, in vitro compartmentalization (IVC), compartmentalized partnered replication (CPR) and phage-assisted continuous directed evolution (PACE). There are some common points between screening and selection strategies. The selected progeny are used as parents in the next round of selection. The process is repeated until the protein with desired properties is achieved.
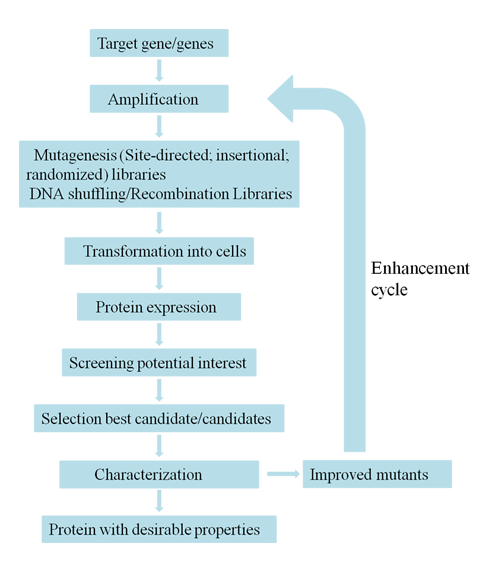
Creative Biostructure offers various tools suitable for every stage of directed evolution, which include diversification library construction, screening-and-selection platform development as well as high-throughput assay development. With advanced technologies and rich experiences in protein engineering, we can perform directed evolution professionally and effectively to meet every customer`s requirements.
Popular strategies used for protein engineering:
- Rational design
- De novo design
- Directed evolution
- Semi-rational design
Ordering Process
Reference:
Michael S. Packer and David R. Liu. Methods for the directed evolution of proteins. Nat Rev Genet. 2015 Jul;16(7):379-94.
