Degenerated Mutagenesis Libraries
Creative Biostructure has developed various approaches for mutagenesis libraries construction. With the advanced mutagenesis libraries construction platform established for a long time, scientists from Creative Biostructure provides unique customized degenerated mutagenesis libraries construction services for you.
Protein libraries that contain defined amino acid mixtures at defined positions are frequently used by protein evolution and engineering applications. Degenerate codon libraries can availably dispose the experimental library-size limitations of directed evolution, they make this achieved via paying attention to diversity of the positions and amino acids which are most possibly to form hits. In addition, they can be formed to cover as little or as much diversity at specially positions as needed, more importantly, they are inexpensive to make. Combining degenerate codons during oligonucleotide synthesis that includes mixtures of nucleotides at each position is a general method to introduce diversity. In most cases, NNK or NNS codons is used by the complete set of standard amino acids, where K = G or T and S = C or G.
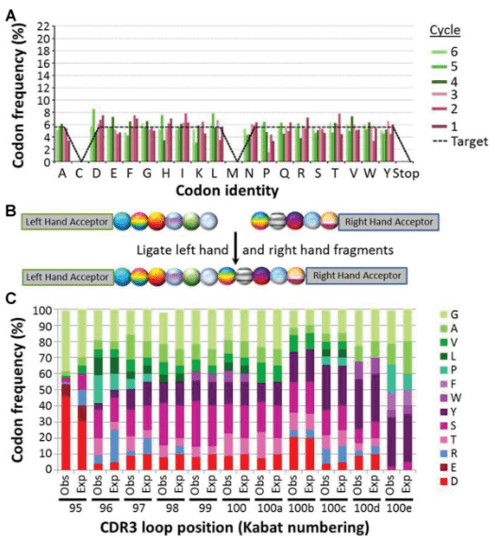 Figure 1. Library construction with controlled codon ratios.
Figure 1. Library construction with controlled codon ratios.
- Strategies for Degenerated Mutagenesis Libraries Construction
Creative Biostructure has developed an algorithm that forms a group of possible degenerate codon libraries, their resulting size, and their score involves in a user-defined scoring function. This library design approach is especially appropriate for coupling computational protein design and multiple sequence alignment data to library construction and screening. Three different methods available for designing library: random sampling of all feasible libraries, scoring of all possible libraries and just scoring libraries within a given size and score constraint.
- Advantages of Degenerated Mutagenesis Libraries
A current variation on degenerate codon libraries is a group of variants, in which all of the 19 amino acid substitutions are enumerated at each position successively. This library is delivered as individually arrayed variants with defined sequences or as a hybrid pool of all other 19 feasibilities for each position. This extremely decrease the screening burden, as well as it has advantages in patent filing for it is possible to test every amino acid mutation in a protein.
Creative Biostructure offers degenerated mutagenesis libraries construction services with library complicacy up to 109 delivered as dsDNA, clones with plasmid vector your chose, or transformants in glycerol stocks.
Creative Biostructure provides various protein engineering and analysis services, including protein evolution. Please feel free to contact us to discuss your project!
Ordering Process
References:
Mena M A, Daugherty P S. Automated design of degenerate codon libraries[J]. Protein Engineering Design and Selection, 2005, 18(12): 559-561.
Jacobs T M, Yumerefendi H, Kuhlman B, et al. SwiftLib: rapid degenerate-codon-library optimization through dynamic programming[J]. Nucleic acids research, 2015, 43(5): e34-e34.
Zheng W, Ye X, Friedman A M, et al. Algorithms for selecting breakpoint locations to optimize diversity in protein engineering by site-directed protein recombination [C]//Proc. CSB. 2007, 6: 31-40.
