4D NMR Services
NMR spectroscopy is a powerful tool for the study of chemical structure using simple one-dimensional techniques. The application of 2D (two-dimension) NMR is used to determine the structure of more complicated molecules since 1980s. 2D-FT (Fourier-transform) NMR can be used in detecting small protein structures within up to 100 amino acids. Over the recent years, multi-dimensional NMR experiments (3D and 4D NMR) have been greatly developed. The major motivation is that standard 2D or 3D NMR experiments cannot resolve the structures of individual compounds contained in complex mixtures; however multi-dimensional (nD) NMR experiments (for example, 4D NMR) is capable of structural characterization of macromolecules such as proteins and complex nucleic acids.
In 2D NMR there is only one systematically varied time period in the sequence of pulses, which modulates the intensity and phase of the detected signals. In 3D NMR, two time periods are varied independently. While in 4D NMR, three time periods are varied. With advanced protein isotope labeling technique, 3D and 4D NMR enable structural elucidation of large proteins of around 15-35 kDa. In 4D NMR spectroscopy, there are three pulses, as the experiment is repeated, and the pulse timings are systematically varied and the oscillations of the spin system are probed point by point in the time domain.
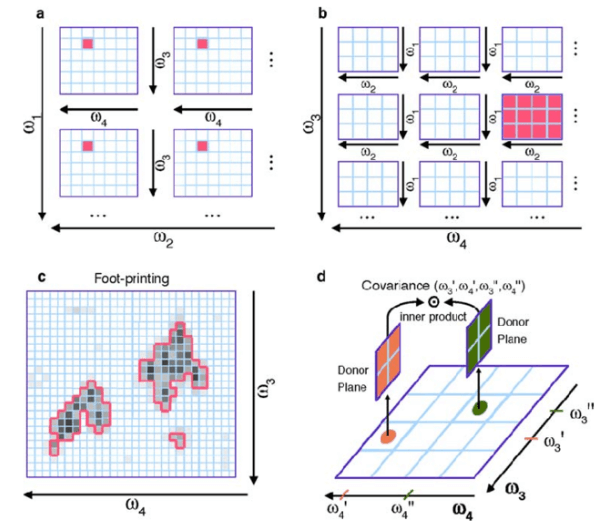 Fig. 1 Representations of a 4D NOESY NMR spectrum.
Fig. 1 Representations of a 4D NOESY NMR spectrum.
At Creative Biostructure, we have state-of-art NMR resources and professional expertise for study of macromolecular structure by using multi-dimensional NMR spectroscopy techniques. 4D spectrum provides unprecedented chemical shift and provides an insight into macromolecular structure, function and dynamics in solution. Creative Biostructure offers custom protein structure determination service based on our in-house NMR platform. We provide one-stop structure identification service, which covers every step from sample preparation to NMR structure elucidation.
Creative Biostructure promises to work closely with our customers to provide excellent services from sample preparation to 4D NMR data collection and analysis. Our goal is to assist our customers to pursue high quality research results.
Please feel free to contact us for a detailed quote.
Ordering Process
References:
- David A. Snyder Æ Fengli Zhang Æ Rafael Bru¨schweiler, Covariance NMR in higher dimensions: application to 4D NOESY spectroscopy of proteins. J Biomol NMR, 2007; 39:165-175.
- Nicholle G. A. Bell, Adam A. L. et. al., Isotope-Filtered 4D NMR Spectroscopy for Structure Determination of Humic Substances. Angew. Chem. Int. Ed. 2015; 54:8382 -8385.
- Nicholle G. A. Bell, Lorna Murray et. al., NMR methodology for complex mixture ‘separation’. Chem. Commun. 2014; 50:1694
