Compound Structure Determination
Structure determination is a cornerstone of modern chemistry and biochemistry, allowing scientists to understand the arrangement of atoms in molecules and their chemical behavior. This process underpins the discovery of new drugs, the development of materials, and advances in fields such as environmental science and molecular biology. Techniques such as Nuclear Magnetic Resonance (NMR) spectroscopy, X-ray crystallography, Mass Spectrometry (MS), and Infrared (IR) Spectroscopy provide precise and complementary information about molecular structures. These methods provide insight into atomic connectivity, functional groups, and three-dimensional arrangements, contributing to applications ranging from pharmacology to nanotechnology.
At Creative Biostructure, we utilize state-of-the-art Nuclear Magnetic Resonance (NMR) platform to provide customized gene-to-structure services for a wide range of biological compounds, including small proteins, peptides, and their complexes. Our comprehensive suite of services includes sample preparation, data acquisition, structure calculation, and molecular modeling, all tailored to meet the specific needs of researchers.
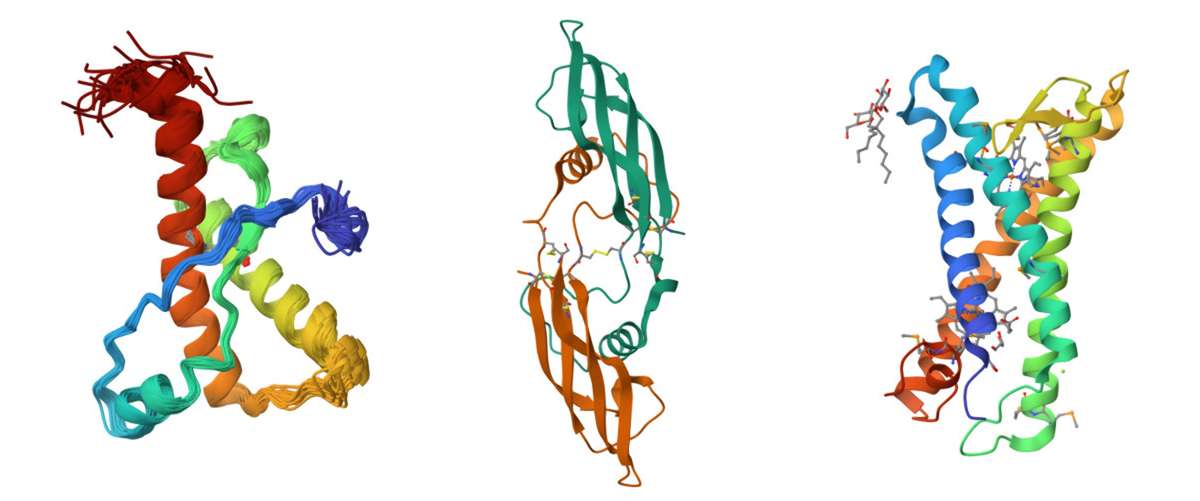 Figure 1. Protein structure examples. Left: Solution structure of the sheep prion protein with polymorphism H168, PDB code: 1XYU. Middle: Crystal structure of human artemin, PDB code: 2GYR. Right: Structure of E. coli superoxide oxidase, PDB code: 5OC0.
Figure 1. Protein structure examples. Left: Solution structure of the sheep prion protein with polymorphism H168, PDB code: 1XYU. Middle: Crystal structure of human artemin, PDB code: 2GYR. Right: Structure of E. coli superoxide oxidase, PDB code: 5OC0.
Techniques to Determine Compound Structure
Nuclear Magnetic Resonance (NMR) Spectroscopy
Nuclear Magnetic Resonance (NMR) spectroscopy is a powerful technique for elucidating molecular structure by studying the interaction of nuclear spins with a magnetic field. It provides detailed information about the chemical environment of atoms, especially hydrogen (¹H) and carbon (¹³C). Chemical shifts, coupling constants, and integration of peaks allow identification of functional groups, stereochemistry, and connectivity. Advanced NMR techniques such as 2D NMR (e.g., correlation spectroscopy-COSY, heteronuclear single quantum coherence-HSQC, and nuclear Overhauser effect spectroscopy-NOESY) further enhance structure determination by revealing spatial relationships and correlations between nuclei.
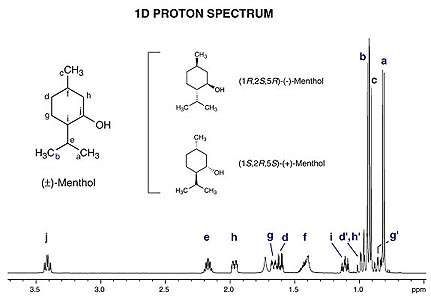 Figure 2. Example ¹ NMR spectrum (1-dimensional) of a mixture of menthol enantiomers plotted as signal intensity (vertical axis) vs. chemical shift (in ppm on the horizontal axis). Signals from spectrum have been assigned hydrogen atom groups (a through j) from the structure shown at upper left.
Figure 2. Example ¹ NMR spectrum (1-dimensional) of a mixture of menthol enantiomers plotted as signal intensity (vertical axis) vs. chemical shift (in ppm on the horizontal axis). Signals from spectrum have been assigned hydrogen atom groups (a through j) from the structure shown at upper left.
NMR spectroscopy relies on multidimensional nuclear magnetic resonance experiments to decode the structural intricacies of biological macromolecules. This process involves several key steps:
Initial Spectrum Analysis
The initial step in NMR-based structure determination is recording a 2D heteronuclear single quantum coherence (HSQC) spectrum. For example, a ¹H, ¹⁵N-HSQC spectrum typically displays one signal for each amino acid residue, except proline. This spectrum serves as a "fingerprint" that provides an overview of the structural integrity of the protein and its feasibility for subsequent experiments.
Resonance Assignment
To analyze NMR data, it is critical to assign resonances to individual nuclei within the protein. This involves performing various 3D NMR experiments, particularly on isotopically labeled proteins (e.g., ¹³C and ¹⁵N double-labeled samples).
Structure Calculation and Validation
Once the resonance assignments and constraints are obtained, the structural model is calculated using specialized computational tools. The process includes structure calculation and structural model validation. Computational programs convert experimental data and known protein properties (e.g., bond lengths and angles) into mathematical energy terms. These programs optimize the structure by minimizing energy states, resulting in an ensemble of models that best fit the experimental data. The accuracy of these models is verified using validation tools. Validation ensures that the final structural model is consistent with experimental data and established biophysical principles, increasing its reliability for downstream applications.
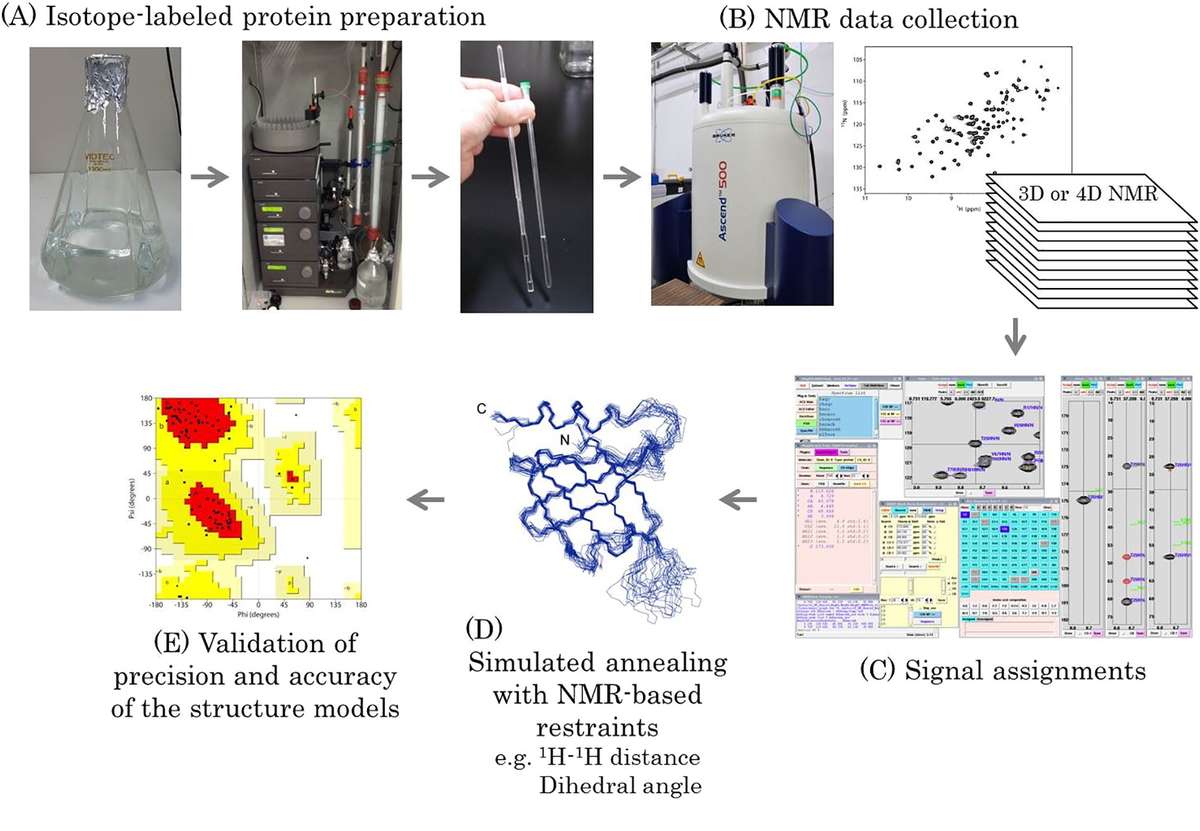 Figure 3. Workflow of protein structure determination by solution NMR spectroscopy. (Sugiki et al., 2017)
Figure 3. Workflow of protein structure determination by solution NMR spectroscopy. (Sugiki et al., 2017)
Mass Spectrometry (MS)
Mass spectrometry determines the molecular mass and fragmentation patterns of compounds, providing clues to molecular formulas and structural elements. Techniques such as Electron Ionization (EI), Matrix-Assisted Laser Desorption Ionization (MALDI), and Tandem MS (MS/MS) allow for detailed characterization of complex molecules. High-resolution mass spectrometry (HRMS) provides accurate mass measurements, aiding in the identification of unknown compounds.
X-ray Crystallography
X-ray crystallography provides three-dimensional structural information by analyzing the diffraction patterns of X-rays passing through a crystal. This technique is indispensable for determining the precise atomic arrangement in solids, including biomolecules like proteins and nucleic acids. Advances in synchrotron radiation and cryo-cooling have improved resolution, enabling the study of large and complex systems.
Infrared (IR) Spectroscopy
IR spectroscopy identifies functional groups within molecules by analyzing their vibrational transitions when exposed to infrared radiation. Characteristic absorption peaks in the IR spectrum correspond to specific bonds, providing insight into molecular composition and structure. Fourier Transform Infrared (FTIR) spectroscopy enhances sensitivity and resolution, making it suitable for both qualitative and quantitative analysis.
Our Comprehensive NMR-Based Services
Creative Biostructure offers complete solutions for the structure determination of compounds by NMR spectroscopy. Our services include:
- Isotope-Labeled Protein Preparation: Efficient production of ¹³C, ¹⁵N, and ²H-labeled samples for high-quality spectra.
→ Isotope Labeled Protein Production - Multidimensional Spectra Collection: Acquisition of advanced 2D, 3D, and 4D NMR spectra tailored to specific project needs.
→ Solid-state NMR Services
→ 4D NMR Services - Data Interpretation: Detailed analysis of spectral data to extract meaningful structural insights.
→ NMR Data Processing and Interpretation - Structure Calculation and Validation: Accurate computational modeling and stringent validation of structural models.
→ Determination of Protein Structure and Dynamics - Molecular Modeling: Creation of high-resolution models for proteins, peptides, and their complexes.
→ Model Reconstruction of Crystal Structures Using Cryo-EM and NMR
At Creative Biostructure, we are committed to advancing structural biology through reliable and efficient NMR structure determination services, including isotope-labeled sample preparation, protein and complex structure determination, and molecular modeling. Please feel free to contact us or submit an online inquiry.
Reference
- Sugiki T, Kobayashi N, Fujiwara T. Modern technologies of solution nuclear magnetic resonance spectroscopy for three-dimensional structure determination of proteins open avenues for life scientists. Computational and Structural Biotechnology Journal. 2017;15:328-339.