High-throughput Screening Using Flow Cytometry
With state-of-art flow cytometry devices and years of experience in the field of high throughput screening, Creative Biostructure now provides customers our most advanced custom high-throughput screening services using flow cytometry methods for protein directed evolution programs as well as other applications.
Flow cytometry is a laser-based biophysical technology that can simultaneously measure and analyze multiple physical characteristics of cells or other single particles. Cells and any other particles from 0.2–150 micrometers in size are suitable for flow cytometry analysis. In a flow cytometry assay, sample cells or particles are carried to a laser intercept in a fluid stream, then suspended and passed by an electronic detection apparatus. More than 5 characteristics of the samples can be determined in just one flow cytometry experiment, including relative size, relative granularity or internal complexity, and relative fluorescence intensity etc. Flow cytometry is also a powerful tool to achieve high-throughput screening (HTS) with advantages of fast, sensitive, cell-by-cell analysis, etc.
Creative Biostructure have developed the most advanced high-throughput screening flow cytometry (HTS-FC) platform for various applications. Our science team can perform both fluorescence-activated cell sorting (FACS)-based and microfluidics HTS process for different protein evolution and engineering applications and your special requirements.
FACS-based high-throughput screening platforms
The FACS stands for fluorescence-activated cell sorting method. FACS-based method relies on non-diffusing fluorescent reporters for the automated identification and isolation of cells that contain different desired gene or protein variants. Cytosolic fluorescent or luminescent proteins are the key for the screening of enzymes such as recombinases, chaperones and inteins by FACS. With the application of fluorescent-labeled antibodies, non-diffusive cell surface-displayed epitopes can also be identified by FACS platforms. Creative Biostructure have developed various antibody fragment display systems to combine with FACS screening and selection methods for high-throughput screening, including yeast display, mammalian cell display, phages display, ribosome display, etc.
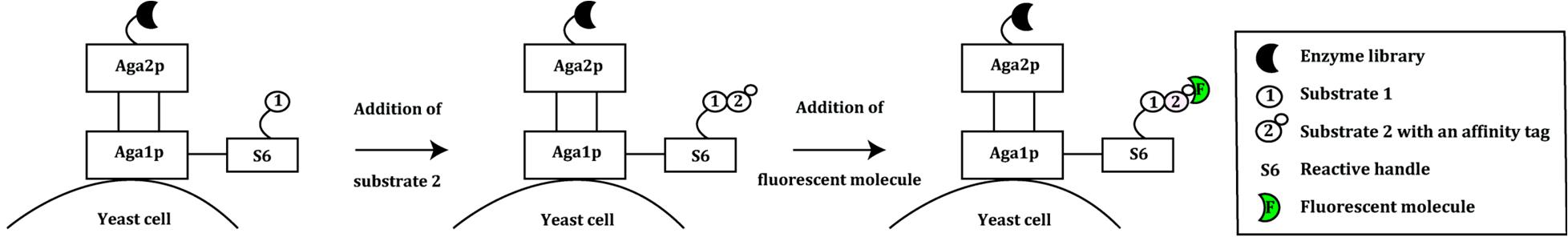
Fig. 1 An overview of the yeast display system used in the evolution of bond-forming enzymes. (Magdalena Wójcik, et al. Int. J. Mol. Sci. 2015, 16, 24918-24945;)
Microfluidics high-throughput screening platforms
Microfluidics is an advanced technique of manipulating and controlling fluids in networks of channels with lowest dimensions from tens to hundreds micrometers. The microfluidics screening platforms have been developed and involved into the field of high-throughput screening because of their advantages such as low sample consumption, easy handling of nanoliter-volumes of liquids, low analysis cost and being suitable for cell-based assays. Microfluidics platforms enable both the droplet manipulation, incubation and sorting in one single system. In Creative Biostructure, different perfusion flow mode, droplet mode, and microarray mode high-throughput screening platforms have been established and are all available for customers to meet different experimental requirements.
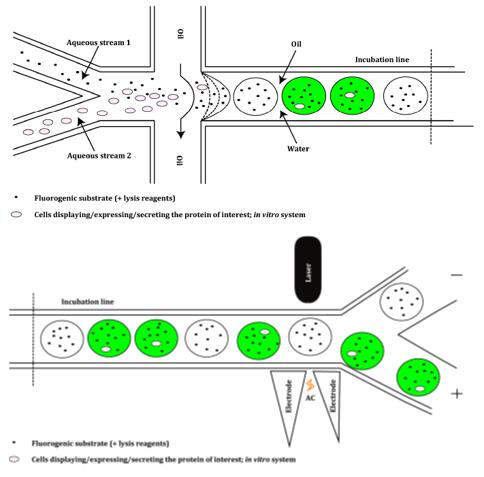 Fig. 2 Representation of a droplet generator module of a microfluidic screening platform. (Magdalena Wójcik, et al. Int. J. Mol. Sci. 2015, 16, 24918-24945;)
Fig. 2 Representation of a droplet generator module of a microfluidic screening platform. (Magdalena Wójcik, et al. Int. J. Mol. Sci. 2015, 16, 24918-24945;)
Creative Biostructure can provide you the perfect flow cytometry based high-throughput screening services for protein engineering and evolution assays as well as other applications.
Welcome to contact us for more details.
References:
Young S M, Bologa C, Prossnitz E R, et al. High-throughput screening with HyperCyt® flow cytometry to detect small molecule formylpeptide receptor ligands[J]. Journal of biomolecular screening, 2005, 10(4): 374-382.
Wójcik M, Telzerow A, Quax W J, et al. High-Throughput Screening in Protein Engineering: Recent Advances and Future Perspectives[J]. International journal of molecular sciences, 2015, 16(10): 24918-24945.
Du G, Fang Q, den Toonder J M J. Microfluidics for cell-based high throughput screening platforms—A review[J]. Analytica chimica acta, 2016, 903: 36-50.