Custom MemPro™ Peptidase MA Services
Creative Biostructure has developed custom MemPro™ gene-to-structure services for Peptidase MA in the protein expression, purification, structure determination and functional analysis.
Metallopeptidases belong to hydrolases, of which the enzymatic mechanism is the nucleophilic attack on a peptide bond mediated by a water molecule. Depending on the number of metalions required for the catalysis, metallopeptidases can be mainly divided into two groups: one group employs only one metal ion for catalysis, while the other group, namely Peptidase MA, needs to coordinate two metal ions. The upper-case letters in the name of a peptidase suggest the catalytic types: “M” represents for metallo-type, A for aspartic-type, S for serine-type, T for threonine-type, C for cysteine-type, and U for unknown type.
Peptidase MA clan is one of the two zinc-dependent metallopeptidases, containing the HEXXH motif with the two His residues as zinc ligands. Generally, a metal ion is coordinated via three ligands. The peptidase MA clan to be further divided into two subclans based on the third ligand types:1) subclan MA(E), also known as the ‘Glu-zincins’, employing a glutamate residue as the third ligand; 2) subclan MA(M), also known as the ‘Met-zincins’, employing histidine or aspartate.
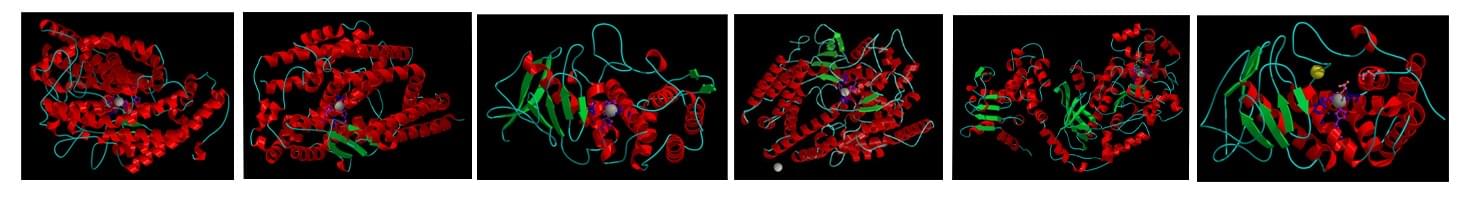
Structures have been solved for several families of the peptidase MA clan, showing the active sites located between the two conserved domains (e.g., the HEXXH motif-containing N-terminal domain is highly conserved among the peptidases). Information of known families in the peptidase MA clan is listed below. Creative Biostructure can also determine the unknown structures of peptidases with/without the inhibitors co-crystallized, based on X-ray crystallography method and known homologs.
Creative Biostructure can provide custom MemPro™ gene-to-structure services for membrane proteins. Click for more information.
| Family |
Family Type Peptidase |
Structure known | Family | Family Type Peptidase | Structure known | Family |
Family Type Peptidase |
Structure known |
| M35 | deuterolysin | Yes | M85 | NleC peptidase | - | M57 | prtB g.p. | - |
| M01 | aminopeptidase N | Yes | M90 | MtfA peptidase | Yes | M64 | IgA peptidase | - |
| M02 | angiotensin-converting enzyme peptidase unit 1 | Yes | >M91 | NleD peptidase | - | M66 | StcE peptidase | - |
| M03 | thimet oligopeptidase | Yes | M93 | BACCAC_01431 g.p. | - | M72 | peptidyl-Asp metallopeptidase | - |
| M04 | thermolysin | Yes | M98 | YghJ g.p. | - | M80 | Wss1 peptidase | - |
| M05 | mycolysin | - | M06 | immune inhibitor A peptidase | - | M84 | MpriBi peptidase | - |
| M09 | bacterial collagenase V | Yes | M07 | snapalysin | Yes | M97 | EcxAB peptidase | - |
| M13 | neprilysin | Yes | M08 | leishmanolysin | Yes | M78 | ImmA peptidase | - |
| M26 | IgA1-specific metallopeptidase | - | M10 | matrix metallopeptidase-1 | Yes | M36 | fungalysin | - |
| M27 | tentoxilysin | Yes | M11 | gametolysin | - | M41 | FtsH peptidase | Yes |
| M30 | hyicolysin | - | M12 | astacin | Yes | M48 | Ste24 peptidase | Yes |
| M32 | carboxypeptidase Taq | Yes | M43 | cytophagalysin | Yes | M49 | dipeptidyl-peptidase III | Yes |
| M34 | anthrax lethal factor | Yes | M54 | archaelysin | - | M56 | BlaR1 peptidase | - |
| M60 | enhancin | - | M61 | glycyl aminopeptidase | - | M76 | Atp23 peptidase | - |